 Some scientific articles are must-reads because of what the scientists accomplished or developed. Some scientific articles are must-reads because of what the scientists discovered. And then some scientific articles are must-reads for both reasons. Here’s an example from a few weeks ago. A group of scientific teams in China worked for years to develop a vehicle that could carry researchers to Hell and back, so they could bring back dirt and animals from Hell. Then they went to Hell, brought back soil, and extensively analyzed the microbial communities in Hell. They discovered that these communities are dramatically distinct from communities in the nearby ocean, and thereby illuminated surprising evolutionary trajectories of the microbial organisms that inhabit Hell.
Some scientific articles are must-reads because of what the scientists accomplished or developed. Some scientific articles are must-reads because of what the scientists discovered. And then some scientific articles are must-reads for both reasons. Here’s an example from a few weeks ago. A group of scientific teams in China worked for years to develop a vehicle that could carry researchers to Hell and back, so they could bring back dirt and animals from Hell. Then they went to Hell, brought back soil, and extensively analyzed the microbial communities in Hell. They discovered that these communities are dramatically distinct from communities in the nearby ocean, and thereby illuminated surprising evolutionary trajectories of the microbial organisms that inhabit Hell.
I’m overdramatizing. The scientists didn’t visit Hell. They visited Hades, which is mythologically different. Okay fine, their inspiring and successful mission went to the hadal zone of the ocean (named, aptly, after the dark underworld of Hades), specifically to the mysterious famous chasms of the Mariana Trench that are nearly 11,000 meters (about 6.8 miles) deep. Somehow, amazingly, microbes can live and reproduce and evolve at that depth. Maybe they’re even happy.
Here is the abstract from their Cell paper. I think it’s a bit too understated, but it is inspiring nonetheless, and we can all use some inspiration right now.
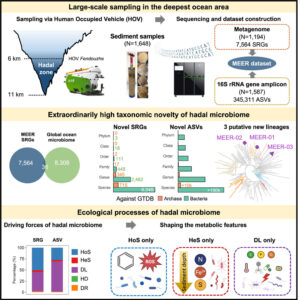
Systematic exploration of the hadal zone, Earth’s deepest oceanic realm, has historically faced technical limitations. Here, we collected 1,648 sediment samples at 6–11 km in the Mariana Trench, Yap Trench, and Philippine Basin for the Mariana Trench Environment and Ecology Research (MEER) project. Metagenomic and 16S rRNA gene amplicon sequencing generated the 92-Tbp MEER dataset, comprising 7,564 species (89.4% unreported), indicating high taxonomic novelty. Unlike in reported environments, neutral drift played a minimal role, while homogeneous selection (HoS, 50.5%) and dispersal limitation (DL, 43.8%) emerged as dominant ecological drivers. HoS favored streamlined genomes with key functions for hadal adaptation, e.g., aromatic compound utilization (oligotrophic adaptation) and antioxidation (high-pressure adaptation). Conversely, DL promoted versatile metabolism with larger genomes. These findings indicated that environmental factors drive the high taxonomic novelty in the hadal zone, advancing our understanding of the ecological mechanisms governing microbial ecosystems in such an extreme oceanic environment.
Microbial ecosystems and ecological driving forces in the deepest ocean sediments
In Cell, 6 March 2025 (this issue of Cell includes three papers and a Commentary piece on the hadal ocean research)
From the groups of Xiang Xiao, Weishu Zhao, Wei-Jia Zhang, Mo Han, Xun Xu, and Shanshan Liu
Snippet by Stephen Matheson
Image credit: graphical abstract from Xiao et al. linked above (CC-BY)